Summary
Background
Mycobacterium tuberculosis whole genome sequencing (WGS) data can provide insights into temporal and geographical trends in resistance acquisition and inform public health interventions. We aimed to use a large clinical collection of M tuberculosis WGS and resistance phenotype data to study how, when, and where resistance was acquired on a global scale.
Methods
We did a retrospective analysis of WGS data. We curated a set of clinical M tuberculosis isolates with high-quality sequencing and culture-based drug susceptibility data (spanning four lineages and 52 countries in Africa, Asia, the Americas, and Europe) using public databases and literature curation. For inclusion, sequence quality criteria and country of origin data were required. We constructed geographical and lineage specific M tuberculosis phylogenies and used Bayesian molecular dating with BEAST, version 1.10.4, to infer the most recent common susceptible ancestor age for 4869 instances of resistance to ten drugs.
Findings
Between Jan 1, 1987, and Sept 12, 2014, of 10 299 M tuberculosis clinical isolates, 8550 were curated, of which 6099 (71%) from 15 countries met criteria for molecular dating. The number of independent resistance acquisition events was lower than the number of resistant isolates across all countries, suggesting ongoing transmission of drug resistance. Ancestral age distributions supported the presence of old resistance, 20 years or more before, in most countries. A consistent order of resistance acquisition was observed globally starting with resistance to isoniazid, but resistance ancestral age varied by country. We found a direct correlation between gross domestic product per capita and resistance age (r2=0·47; p=0·014). Amplification of fluoroquinolone and second-line injectable resistance among multidrug-resistant isolates is estimated to have occurred very recently (median ancestral age 4·7 years [IQR 1·9–9·8] before sample collection). We found the sensitivity of commercial molecular diagnostics for second-line resistance to vary significantly by country (p<0·0003).
Interpretation
Our results highlight that both resistance transmission and amplification are contributing to disease burden globally but vary by country. The observation that wealthier nations are more likely to have old resistance (most recent common susceptible ancestor >20 years before isolation) suggests that programmatic improvements can reduce resistance amplification, but that fit resistant strains can circulate for decades subsequently implies the need for continued surveillance.
Funding
National Institutes of Health, a Harvard Institute of Global Health Burke Fellowship from the National Human Genome Research Institute, National Institutes of Health, the Bill & Melinda Gates Foundation, Boston Children’s Hospital, Bushrod H Campbell and Adah F Hall Charity Fund, and Charles A King Trust Postdoctoral Fellowship.
Introduction
The emergence of multidrug-resistant and extensively drug-resistant tuberculosis presents a major obstacle to efforts to accelerate tuberculosis decline. Halting the transmission of drug-resistant tuberculosis has been a major focus of studies addressing the emergence of drug-resistant tuberculosis.
But the epidemic is ultimately defined by local factors that remain understudied in many parts of the world.
The study of geographical and temporal heterogeneity of the drug-resistant tuberculosis epidemic can provide insights into these local factors as key drivers of multidrug-resistant tuberculosis prevalence and persistence in the community, including programmatic and bacterial factors. This understanding is key to future disease control and prevention of antibiotic resistance development.
,
,
,
The application of WGS has allowed us to better understand the genetic determinants of drug resistance within M tuberculosis.
The detection of these genetic determinants using molecular technologies that include WGS is now increasingly adopted for tuberculosis resistance diagnosis in many parts of the world
and is beginning to replace the more biohazardous and time consuming culture-based drug susceptibility tests. The study of isolates sampled from epidemiological outbreaks or from the same host over time has allowed the estimation of M tuberculosis‘ molecular clock rate, or temporal rate of accumulation of fixed genome-level variation.
,
The application of this rate to new WGS data from isolates collected for surveillance has helped improve transmission inference and molecular dating of specific evolutionary events such as resistance acquisition or lineage divergence.
,
,
Evidence before this study
Acquisition and spread of drug-resistance by Mycobacterium tuberculosis varies across countries. Local factors driving evolution of drug resistance in M tuberculosis are not well studied. We searched PubMed for the following MeSH terms: “tuberculosis”, “antibacterial drug resistance”, “communicable disease transmission”, and “clonal evolution” with no date restrictions up until Jan 5, 2021. The search yielded no results, which suggested that local factors driving evolution of drug resistance in M tuberculosis are not well studied.
Added value of this study
We applied molecular dating to 6099 global M tuberculosis patient isolates and found the order of resistance acquisition to be consistent across the countries examined (ie, acquisition of isoniazid resistance first followed by rifampicin and streptomycin followed by resistance to other drugs). In all countries with data available, there was evidence for transmission of resistant strains from patient to patient and in the majority for extended periods of time (>20 years). Countries with lower gross wealth indices were found to have more recent resistance to the drug rifampicin. Based on the resistance patterns identified in our study, we estimate that commercial diagnostic tests vary considerably in sensitivity for second-line resistance diagnosis by country.
Implications of all the available evidence
The longevity of resistant M tuberculosis in many parts of the world emphasises its fitness for transmission and its continued threat to public health. The association between country wealth and recent resistance acquisition emphasises the need for continued investment in tuberculosis care delivery and surveillance programmes. Geographically relevant diagnostics that take into account a country’s unique distribution of resistance are necessary.
We aimed to use a large clinical collection of M tuberculosis WGS and resistance phenotype data to study how, when, and where resistance was acquired on a global scale. We also assessed the distribution of unexplained M tuberculosis phenotypic resistance across 20 countries, to evaluate the accuracy and geographical heterogeneity of molecular detection of common M tuberculosis genetic resistance determinants, and discuss implications for drug-resistant tuberculosis control.
Results

Figure 1Study profile
The process of identification and exclusion of genomic data included in the study is shown. M tuberculosis=Mycobacterium tuberculosis. WGS=whole genome sequencing.

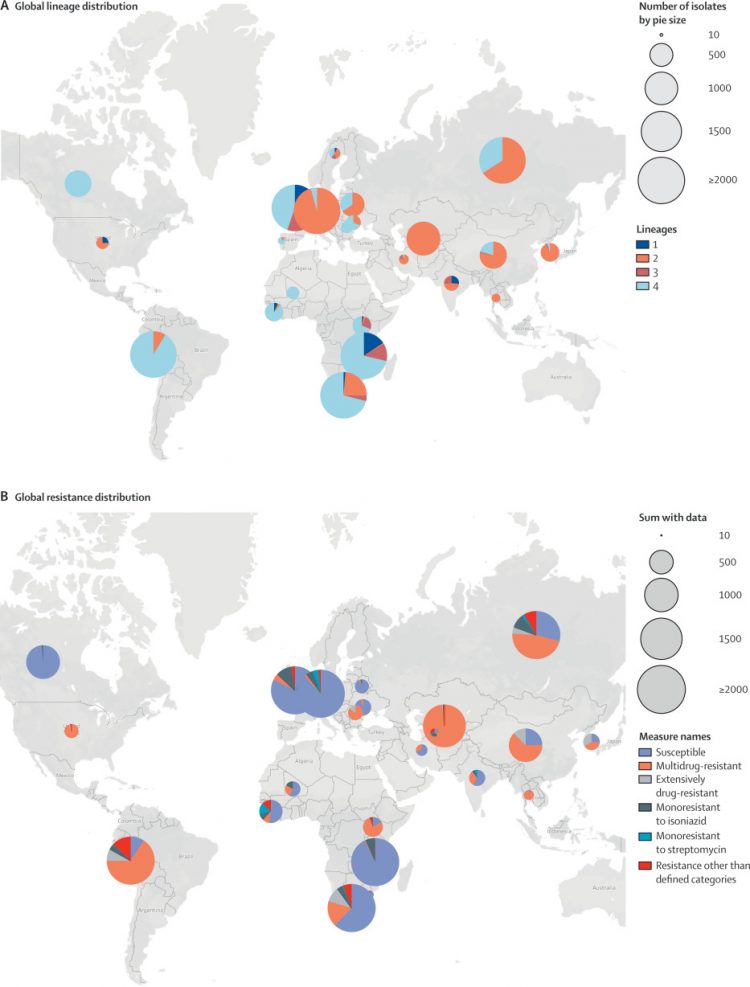
Figure 2Global distribution of Mycobacterium tuberculosis in the study sample
was higher in 14 (67%) countries, and lower in three (14%) countries (appendix p 13). Multidrug resistance rates by lineage were 3% (n=439) for lineage 1, 48% (n=1085) for lineage 2, 4% (n=760) for lineage 3, and 23% (n=3358) for lineage 4.

Figure 3MRSCA distribution by drug (n=4844) and pairwise Wilcoxon rank sum tests comparing MRSCA ages by drug category
Boxplots showing range of MRSCA distribution globally for nine tuberculosis drugs (A) and pairwise Wilcoxon rank sum tests comparing MRSCA ages by drug category (B). Red indicates p<0·001 (Bonferroni threshold); pink indicates p<0·01; and white indicates p≥0·01. MRSCA=most recent susceptible common ancestor.

Figure 4Proportion of multidrug-resistant isolates with recent amplification of resistance
Resistance to ethambutol, pyrazinamide, fluoroquinolones, or second-line injectables by country (MRSCA age estimate <5 years ago) are shown. The key lists the number of multidrug-resistant isolates analysed from each country. Error bars indicate 95% CI. Four countries displayed a measurable proportion of recent fluoroquinolone and second-line injectable amplification (95% CI does not include 0): China, Peru, Romania, and South Africa. MRSCA=most recent susceptible common ancestor.

Figure 5MRSCA distribution per country and median rifamycin MRSCA date versus GDP per capita for 12 countries
and their geographical distribution among the 8550 isolates with country of origin and WGS data meeting quality criteria (figure 1). We pooled results across lineages in each country. Resistance mutation prevalence varied significantly by country. The most frequent isoniazid causing mutation,
katG Ser315Thr, was more frequent among phenotypically isoniazid-resistant isolates by drug susceptibility tests (pheno-R) in Russia (444 [84%] of 526) than in Peru (510 [67%] of 760; Fisher’s exact test pfabG1-inhA promoter, was more prevalent among isoniazid pheno-R Peruvian isolates (20%) than in Russian isolates (8%; Fisher’s exact test pfabG/inhA promoter −15 cytosine to thymine (appendix p 31). The mutation Ile491Phe was described in 2017 to be common in Eswatini
and is not detectable by Hain Line-probe or GeneXpert commercial molecular diagnostics. In our sample that did not contain data from Eswatini, we calculated an SD of 1% (range 0–4) for the global frequency of Ile491Phe among rifamycin pheno-R isolates.
TableSensitivity and specificity of commercial and WGS-based tests for resistance diagnosis
Sensitivity is the percentage of resistant isolates classified as resistant. Specificity is the percentage of susceptible isolates classified as susceptible. WGS=whole genome sequencing.
or randomForests
might improve sensitivity and specificity. We found this select-WGS approach to improve sensitivity slightly for isoniazid and second-line injectables with relatively preserved specificity (table). In addition, this approach allowed prediction for drugs not tested by commercial diagnostics—pyrazinamide, ethambutol, and streptomycin. For comparison, we assessed if including any non-silent variant in the resistance regions (excluding a select number of known lineage markers) was indeed inferior to the more informed select WGS test reported previously. We found that the all WGS test only modestly improved sensitivity and at the expense of a larger decrease in specificity (table).
Discussion
We found multidrug-resistant tuberculosis in nearly every country represented by more than ten isolates. Extensively drug-resistant isolates were found in half of these countries and spanned all five major continents. Lineage 2 had the highest percentage of multidrug-resistant isolates in our sample followed by lineages 4, 3, and 1. Using this diverse sample, we dated more than 4869 resistance phenotypes across four lineages and 15 countries.
Our results corroborate these findings using phenotypic resistance data across a larger, geographically diverse sample. After isoniazid, we found that M tuberculosis acquires resistance to rifamycins or streptomycin, then ethambutol, followed by pyrazinamide, ethionamide–prothionamide, fluoroquinolones, second-line injectables, or cycloserine. We found no correlation between the age of resistance acquisition and the year of clinical introduction of the drug but there might be multiple other causes for the observed order of resistance acquisition. Differences in mutation rates across drug targets or resistance genes have been postulated but shown to be an unlikely explanation for isoniazid resistance arising first.
Pharmacokinetic differences might result in higher risk for underdosing for some drugs and earlier resistance acquisition.
Bacterial fitness costs are also variable across resistance mutations. For isoniazid resistance, mutations like katG Ser315Thr carry a low fitness cost and probably contribute to resistance arising earliest for this drug.
The order of drug administration can explain dating differences between first-line (isoniazid, rifamycins, ethambutol, pyrazinamide) and second-line (ethionamide–prothionamide, fluoroquinolones, second-line injectables) or third-line (cycloserine) resistance, as second-line drugs are usually only administered after resistance to first-line drugs is ascertained. Acquisition of resistance to isoniazid first and then rifamycins might also relate to their use for treatment of latent tuberculosis infection, leading to more exposure and selection pressure overall. However, because adoption of isoniazid preventive therapy for latent tuberculosis remains low in many parts of the world, we expect it to be a lesser contributor to isoniazid and rifamycin resistance.
Lastly, the observation of contemporaneous acquisition of rifamycin and streptomycin resistance is probably best explained by the effects of category II tuberculosis treatment initially recommended in 1991.
Category II is no longer recommended by WHO but consists of adding streptomycin to the first-line drug regimen after treatment failure. Our dating supports that streptomycin resistance amplified among patients whose treatment was unsuccessful due to recent rifamycin resistance or multidrug-resistant acquisition, or both.
Thus, the identification of isolates with old resistance in most countries with available data suggests high fitness for continued transmission between human hosts. This hypothesis is also supported by the finding of a lower bound of tuberculosis resistance due to transmission at 14–52% across countries with available data. As our approach cannot distinguish between resistance importation through human migration after transmission outside of the country and new resistance acquisition, these figures are underestimates of the true resistance burden due to transmission. Tuberculosis mathematical models have previously predicted transmission to be a major driver of observed resistance rates and have emphasised that drug resistant strain fitness is a key parameter dictating how the resistance epidemic will unfold.
Our results support that some resistant isolates are fit and successfully transmitting patient-to-patient, sometimes uninterrupted, for more than 20 years. These data emphasise the need to contain resistance transmission through improved diagnosis, treatment, and other preventive strategies such as infection control and vaccine development.
We estimate that half of fluoroquinolones and second-line injectable resistant isolates had acquired resistance within 4·7 years of isolation despite the promotion of directly observed therapy by WHO since 1994. As most fluoroquinolones and second-line injectable resistant isolates are also multidrug-resistant, our results also emphasise the need for better regimens to treat multidrug resistance that can prevent resistance amplification. By country, we found a significant correlation between the estimated age of resistance acquisition and per capita gross domestic product, with more affluent countries having older ages of resistance. This correlation is likely to be driven by a combination of factors, but the routine use of drug susceptibility tests and close patient monitoring in well resourced health systems are probably important contributors. Specifically, we found that China and the UK had the oldest resistance ages across the drugs. The Chinese national tuberculosis programme budget was increased from US$98 million in 2002 to $272 million in 2007
and a new policy for free tuberculosis diagnostic tests and drug use was introduced in 2004.
This increased investment can explain the observed low rates of recent resistance acquisition in China.
we measured improvements in resistance sensitivity offered by including mutations outside of regions targeted by commercial diagnostics through direct association. This approach offered modest improvements in sensitivity with little to no change in specificity. We found a considerable number of indeterminate mutations in resistance regions, which when included with simple direct association improve sensitivity, but at the expense of loss of specificity. The study of these variants through statistical models will probably further inform their diagnostic use in the future.
Another limitation of our study is the scarcity of data for immigration or disease importation. For this reason, we avoided drawing conclusions concerning specific countries, and instead we comment on trends spanning multiple countries expected to have a range of tuberculosis burden due to importation. The absence of these data also limited our ability to assess disease transmission across borders. Our analysis also assumed accuracy of culture-based phenotypic drug susceptibility tests, even though test to test variability is known to exist. We justify this assumption as our data was curated from ReseqTB,
and studies in which phenotypic testing was done in national or supranational laboratories with rigorous quality control.
YE did the data analysis and drafted and revised the manuscript. LF and AD contributed to the data analysis. MRF conceptualised the study, supervised the data analysis, and reviewed, wrote, and edited the manuscript. YE and MRF verified the data. All authors provided key edits to the manuscript.
We declare no competing interests.