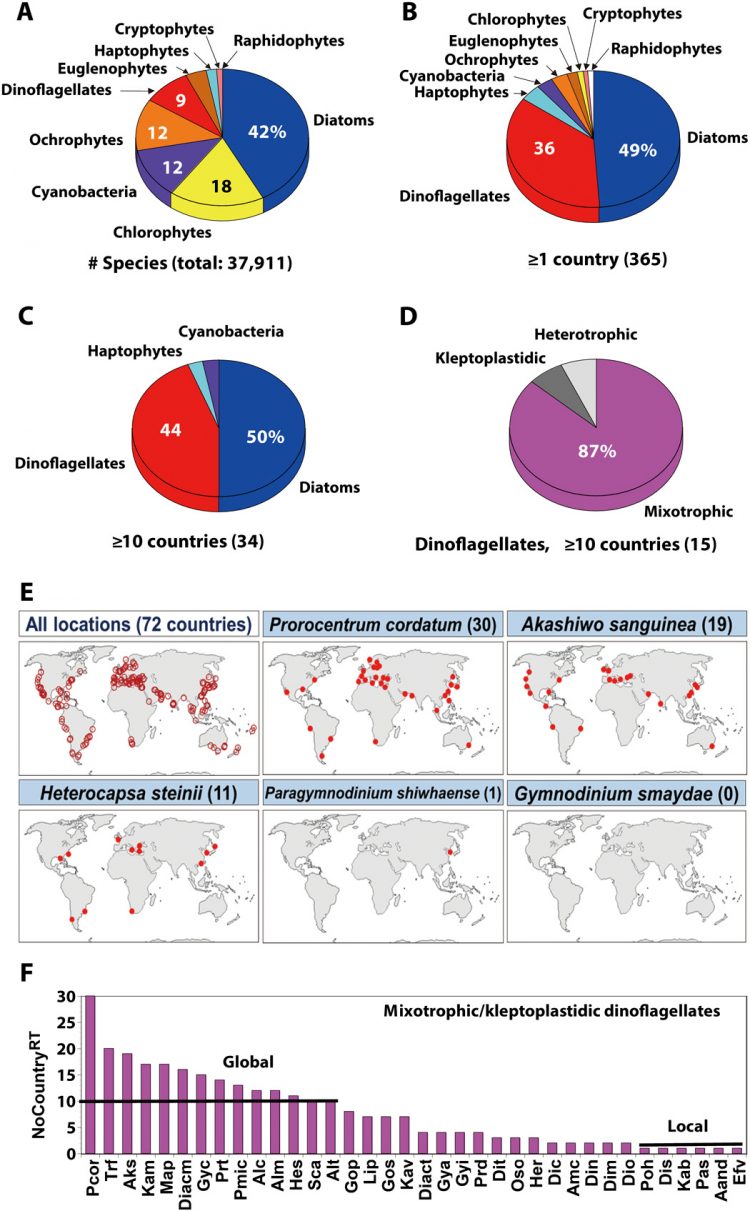
Data on the number of species belonging to each microalgal group
We analyzed the species belonging to each group of the formally described microalgae and cyanobacteria. These data were obtained from Algaebase (https://www.algaebase.org) in March 2020. Only currently accepted species were included. The categorized microalgal groups were diatoms, dinoflagellates, haptophytes, ochrophytes including raphidophytes, chlorophytes, cryptophytes, and euglenophytes. However, raphidophytes were separated from ochrophytes because some raphidophyte species caused red tides globally, whereas there were no ochrophytes, except raphidophytes, that caused red tides globally.
Data on the red tides in the world ocean in 1990–2019
We analyzed red tides occurring in the world ocean during 1990–2019 that had been reported in the literature (~800 references; e.g., tables S2 and S3 for the target mixotrophic dinoflagellates). Furthermore, we investigated the causative species of each red tide event and also the country where the red tide event occurred and, lastly, determined the number of the countries in which each red tide species had caused red tides (NoCountryRT).
Data on presence and absence of photosynthesis-related genes in dinoflagellates
To investigate 22 target genes related to photosynthesis (8 photosystem genes, 9 Calvin cycle genes, and 5 gluconeogenesis genes) of major dinoflagellates, the transcriptomes of 3 heterotrophic, 3 kleptoplastidic, 9 mixotrophic, and 2 autotrophic dinoflagellates were analyzed (Fig. 2 and tables S4 to S10). The transcriptomes of the heterotrophic dinoflagellate Polykrikos kofoidii, the kleptoplastidic dinoflagellate G. smaydae, the mixotrophic dinoflagellates P. shiwhaense and Biecheleria cincta, and the autotrophic dinoflagellate Biecheleriopsis adriatica were newly assembled in this study, but those of the other dinoflagellates were obtained from the literature (table S6). The transcriptome of Heterocapsa rotundata was also analyzed to test whether the signal of the presence of photosynthesis-related genes of G. smaydae was this predator’s own signal or from its prey. On the basis of the assembled transcriptomes, the target genes of the dinoflagellate species were identified using a tBLASTn algorithm as implemented in CLC Genomic Workbench ver. 10.0.1 (QIAGEN N.V. Venlo, the Netherlands) (table S7). Some genes of which phylogenetic relationships need to be confirmed were aligned, and trees were constructed. Furthermore, the presence of psaA and psbB genes of the dinoflagellates were additionally confirmed on the basis of genomic DNA sequencing (figs. S3 and tables S5 and S8 to S10).
Culturing, sequencing, and sequence assembly of six dinoflagellates. Before the transcriptome experiments were conducted, two consecutive single-cell isolations of cells from each clonal culture of P. kofoidii (PKJH1607), G. smaydae (GSSH1005), P. shiwhaense (PSSH0605), B. cincta (BCSW0906), B. adriatica (BATY06), and H. rotundata (HRSH1201) were performed to confirm no potential contamination by bacteria or other small eukaryotes. Furthermore, to confirm rapid growth condition and no remaining prey cells in each culture, 5 ml of aliquots was taken from each bottle every 2 days and fixed with Lugol’s solution (final concentration, 5%). The aliquots were taken from the fixed sample and then transferred to two 1-ml Sedgwick-Rafter chambers for cell enumeration.
For transcriptome analysis, a dense culture (~80 cells ml−1) of P. kofoidii growing on Alexandrium minutum (CCMP1888) was transferred to an 800-ml culture flask containing dense prey (~6000 cells ml−1) and autoclaved filtered seawater. After prey cells were undetected in the ambient waters (2 days after inoculation), P. kofoidii cells were maintained without added prey cells for 3 days (starved for 3 days). For harvesting P. kofoidii cells, 800 ml of aliquot containing approximately 168,000 cells was taken from the culture flask and then centrifuged for 5 min at 800g using a Vision Centrifuge VS-5500 (Vision Scientific Company, Bucheon, Korea). Similarly, a dense culture (2000 cells ml−1) of G. smaydae growing on H. rotundata (HRSH1201) was transferred to a 2-liter polycarbonate (PC) bottle containing dense prey (~60,000 cells ml−1). After prey cells were undetectable (3 days after inoculation), G. smaydae cells were maintained without added prey cells for 3 days. For harvesting cells, 1.8 liter of aliquot containing approximately 4 × 107 cells was taken from the PC bottle and then centrifuged for 10 min at 1000g.
A dense culture (~3000 cells ml−1) of P. shiwhaense growing on Amphidinium carterae (SIO PY-1) was distributed to an 800-ml culture flask containing dense prey (~5000 cells ml−1). After prey cells were undetectable (2 days after inoculation), P. shiwhaense cells were maintained without added prey cells for 18 days. For harvesting cells, 800 ml of aliquot containing approximately 5 × 106 cells was taken from the culture flask and then centrifuged for 10 min at 1000g. Moreover, a dense culture (~3000 cells ml−1) of B. cincta growing on the raphidophyte Heterosigma akashiwo (HAKS9905) was transferred to a 2-liter PC bottle containing dense prey (~25,000 cells ml−1). After a prey cell was undetected in the ambient waters (4 days after inoculation), B. cincta cells were maintained without added prey cells for 3 days. For harvesting cells, 1.8 liter of aliquot containing approximately 2 × 107 cells was taken from the PC bottle and then centrifuged for 10 min at 1000g.
A dense culture of B. adriatica growing autotrophically (~5000 cells ml−1) was distributed to a 2-liter PC bottle containing an autoclaved f/2-Si medium (25). For harvesting cells, 500 ml of aliquot containing approximately 5 × 107 cells in its exponential phase (10 days after inoculation) was taken from the PC bottle and then centrifuged for 10 min at 1000g. Similarly, a dense culture (~10,000 cells ml−1) of H. rotundata growing autotrophically was transferred to an 800-ml culture flask containing an autoclaved f/2-Si medium. Five hundred milliliters of aliquot containing approximately 1 × 107 cells in its exponential phase (7 days after inoculation) was taken from the flask and then centrifuged for 10 min at 1000g.
The pellets of the six dinoflagellate samples harvested were immediately frozen with liquid nitrogen and stored at −80°C until RNA extraction. Then, total RNA from each sample was extracted according to the RNeasy Plant Mini Kit protocol (catalog no. 74903; Qiagen, Germany) and treated with the RNase-Free DNase set (catalog no. 79254) to remove any residual genomic DNA. The complementary DNA (cDNA) libraries of G. smaydae, B. cincta, and B. adriatica were sequenced using a HiSeq 2500 system (Illumina Inc., San Diego, CA) by the National Instrumentation Center for Environmental Management (Seoul, Korea). P. kofoidii, P. shiwhaense, and H. rotundata were sequenced using a NovaSeq 6000 system (Illumina Inc., San Diego, CA) by Macrogen (Seoul, Korea). Moreover, the quality of the data used for each assembly was verified using FastQC v.11.6 (26). Subsequently, the clean reads of each dinoflagellate species were independently de novo assembled with Trinity software (27). The transcriptomes of the six dinoflagellates analyzed in this study were deposited in the National Center for Biotechnology Information (NCBI) Sequence Read Archive (SRA accession numbers SRR11946747, SRR11947552, SRR11994189, SRR11994191, SRR11994206, and SRR12020522).
The data on the transcriptome assembly of the kleptoplastidic dinoflagellate P. piscicida and mixotrophic dinoflagellates Y. yeosuensis and Ansanella granifera were obtained from our previous studies (28–30). Moreover, the transcriptomic sequences of the heterotrophic dinoflagellates Oxyrrhis marina, Noctiluca scintillans, kleptoplastidic dinoflagellate D. acuminata, mixotrophic dinoflagellates Lingulodinium polyedra, Gymnodinium catenatum, A. andersonii, Heterocapsa steinii, and the autotrophic dinoflagellate Pelagodinium bei were obtained from the Marine Microbial Eukaryote Transcriptome Sequencing Project (table S6) (31, 32).
Gene identification. The presence or absence and transcript sequences of target genes encoding for plastid-related proteins in these dinoflagellates was identified using a tBLASTn algorithm as implemented in CLC Genomic Workbench ver. 10.0.1 (QIAGEN N.V. Venlo, the Netherlands). We used a stringent E-value cutoff criterion of E-20. The amino acid sequences of the previously well-identified plastid genes were used as queries to perform tBLASTn searches (table S7). Among the plastid genes belonging to photosystem I and photosystem II, only the genes commonly identified on minicircles of dinoflagellates were analyzed in this study (Fig. 2 and table S4) (33, 34). Regarding query sequences of the psbI gene, however, identifying its orthologous genes of dinoflagellates was impossible due to its small sequence size (approximately 35 to 38 amino acids). Moreover, the possible presence of prey-originated plastid gene sequences in the transcriptome of G. smaydae was further confirmed using the strict criteria of the BLASTn algorithm (cutoff E-value <E-100 and identity >99%) against the transcriptome of the prey H. rotundata. Similarly, the identified plastid genes of the other dinoflagellates grown heterotrophically or kleptoplastically also needed to be analyzed to determine whether they are potentially evolutionarily remnant genes or just remained genes from the prey materials. However, there have been no data about the clonal strain of the prey transcriptomes, except for G. smaydae, and thus, we carried out additional homology searches for these genes against the NCBI nonredundant database. If the homology of the gene was highly similar (i.e., cutoff E-value <E-100 and identity >95%) to that of any species in the genus to which the prey species belongs, then we considered this gene as a prey-originated gene and did not include it in the heatmap. Moreover, these relationships were further validated on the basis of phylogenetic analysis (see the next section).
Phylogenetic analysis for gene validation. Some genes that were present in some dinoflagellates but absent in others were aligned with multiple sequences by MEGA v.4 (35). The alignments of the tBLASTn hits were manually inspected and curated to remove problematic sequences (i.e., chimeric sequences and/or contaminant sequences), and the ambiguously aligned sites were further removed. In this study, the nucleotide sequence–based phylogenies were constructed to eliminate the possibility that fragmented sequences of potential genes are filtered through the decoding process. The phylogenetic analyses were performed under the GTR+G model and inferred by Bayesian analysis using the MrBayes v.3.1 program (36). Bayesian analysis was sampled every 200 generations and continued until the average SD of the split frequencies dropped below 0.01. Moreover, it was confirmed that the analyses reached statistical stationarity well before the burn-in period by plotting the ln-likelihood of the sampled trees against generation time.
Genomic DNA sequencing for gene validation. Since the coding region of plastid genes that we analyzed in this study consisted of a single exon without internal introns, we confirmed the presence of a few identified transcripts (i.e., cDNA of psaA and psbB) by genomic DNA sequencing. Especially, since all the genes in the photosystem identified from the transcriptome of G. smaydae were identical to those of its prey H. rotundata (i.e., no possession of its own genes), we confirmed whether these genes existed inside G. smaydae cells until the cells were almost dead (after 10-day starvation). Thus, we designed the universal polymerase chain reaction (PCR) primers for partial sequences of psaA and psbB genes of the dinoflagellate species listed in tables S8 to S10. To determine the universal sequences, manual searches of the alignments were conducted using the program MEGA v.4. The sequences for the forward and reverse primers for psaA and psbB genes were selected from the regions that are conserved from all the aligned dinoflagellate species (table S10). The primer sequences were analyzed with Primer 3 (Whitehead Institute and Howard Hughes Medical Institute, MD) and Oligo Calc: Oligonucleotide Properties Calculator (37) for optimal melting temperature and secondary structure.
For PCR amplification, the genomic DNAs of some target dinoflagellate species (i.e., 2-day starved G. smaydae, 10-day starved G. smaydae, 5-day starved Y. yeosuensis, and autotrophically growing A. carterae, H. rotundata, and G. catenatum) were extracted using the AccuPrep Genomic DNA Extraction Kit (Bioneer, Daejeon, Korea), according to the manufacturer’s instructions. The PCR conditions were as follows: initial denaturation at 95°C for 2 min; followed by 35 cycles at 95°C for 20 s, an appropriate annealing temperature for 40 s, and 72°C for 1 min, with a final elongation step at 72°C for 5 min. The annealing temperature was adjusted for specific primer sets according to the manufacturer’s instructions. The detailed methods for PCR amplification, sequencing, and alignment were according to the procedures used by Jang et al. (38). If the PCR product mixed with 0.5 μl of goRed fluorescent reagent (Genepole, Seoul, Korea) was not identified from the first amplification as checked using gel electrophoresis, then the second DNA amplification using the same primer sets was performed with the 1 μl of the first PCR product as a template. As a result, the presence or absence of psaA and psbB genes identified from transcriptomic data could be verified by sequencing genomic DNA of partial 400 to 500 lengths of psaA and psbB genes (table S5).
Data acquisition for the calculation of two mixotrophic ability indices
We developed two new indices of mixotrophic ability of a mixotrophic dinoflagellate—predation contribution to total growth rate (PredCTGR) and the ratio of the number of edible prey taxa to that of total tested prey taxa (RETPREY). We selected mixotrophic dinoflagellates of which both autotrophic (without added prey, GRAuto) and total or mixotrophic growth rates (with added prey, GRTotal) had been reported (table S11).
The PredCTGR of a mixotrophic dinoflagellate under a given prey concentration, temperature, and light condition was calculated as follows
We did not calculate PredCTGR when both GRTotal and GRAuto were negative. Furthermore, we gave the PredCTGR value of 100% when GRAuto was zero or negative, whereas GRTotal was positive. In a rare case, GRAuto was slightly greater than GRTotal. We gave the PredCTGR value of 0% in this case.
We calculated the ratio of the number of edible prey taxa to that of total tested prey taxa (RETPREY) of a mixotrophic dinoflagellate rather than the absolute number of edible prey because the prey species for one mixotrophic dinoflagellate species were sometimes different from those of the other mixotrophic dinoflagellate species in different literature. In this calculation, we included the mixotrophic dinoflagellate species for which the number of total tested prey taxa was ≥5 species. Engulfment-feeding mixotrophic dinoflagellates usually do not feed on prey larger than themselves, and thus, we excluded the prey species larger than themselves from the nominator (i.e., total tested prey taxa) when the target mixotrophic dinoflagellates were engulfment feeders.